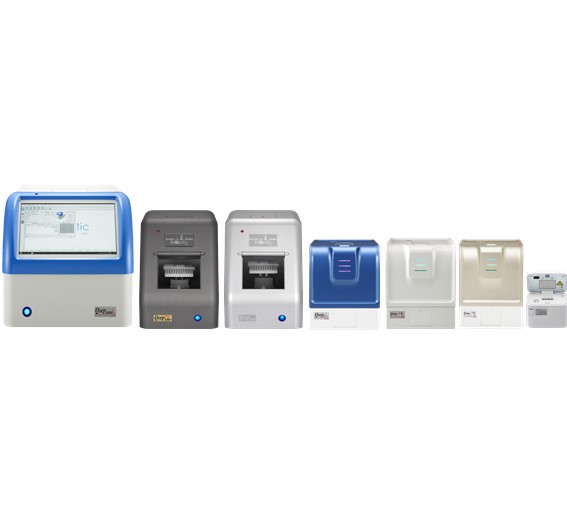
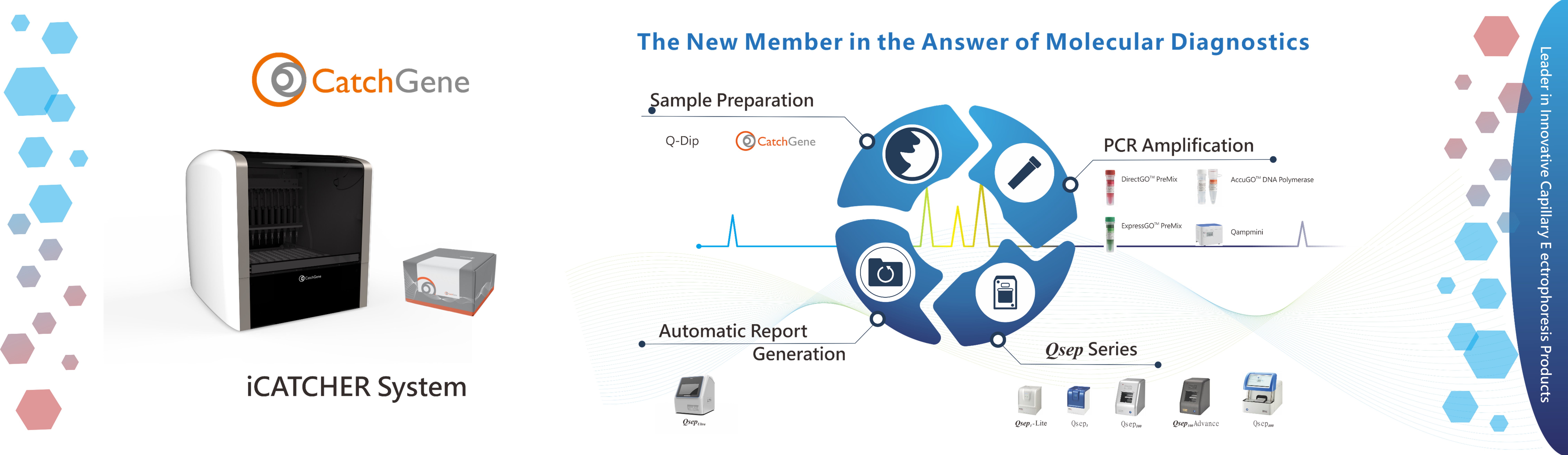
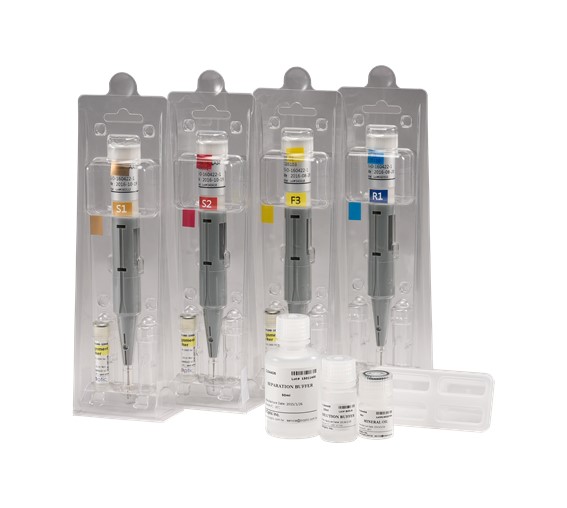
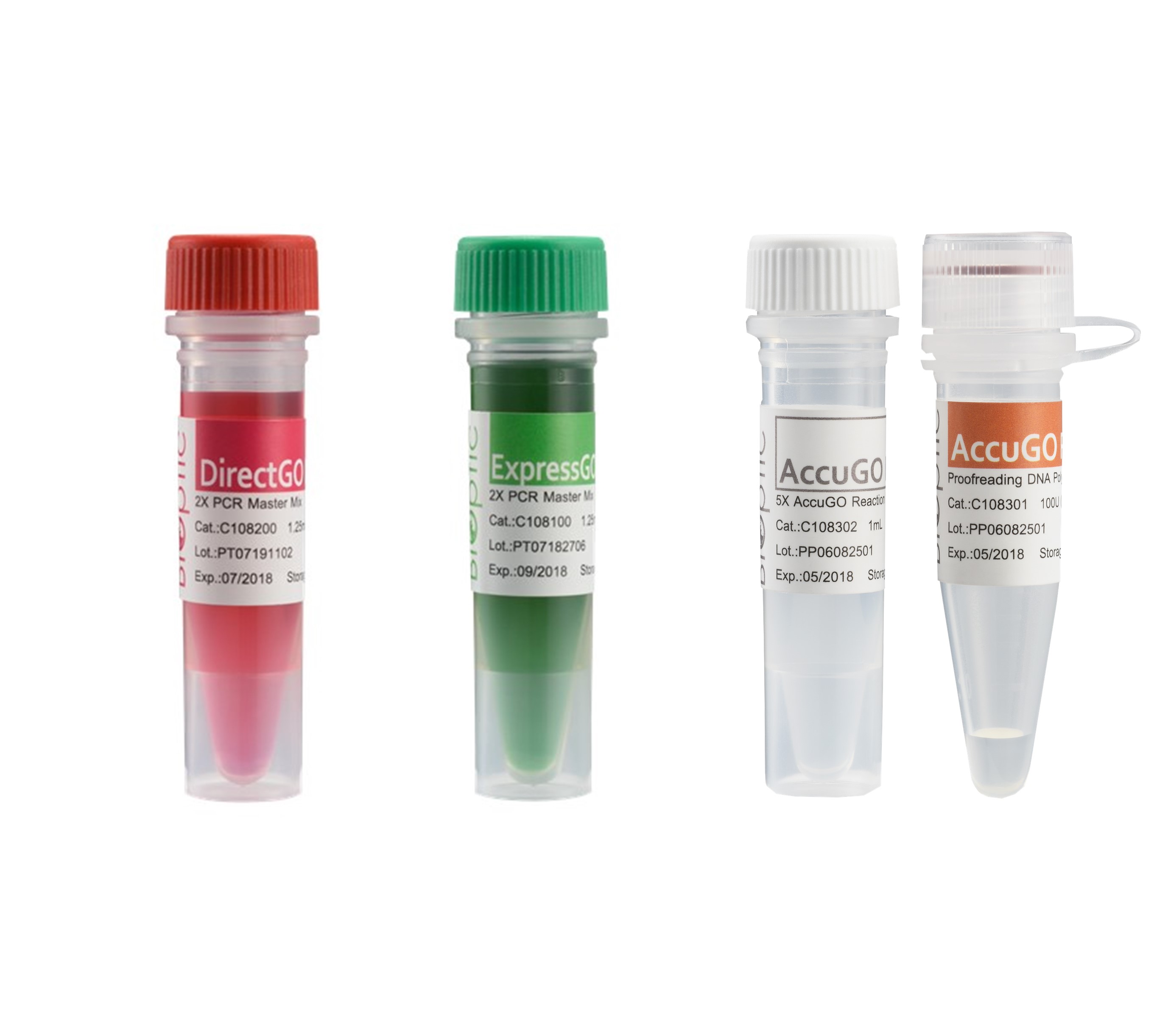
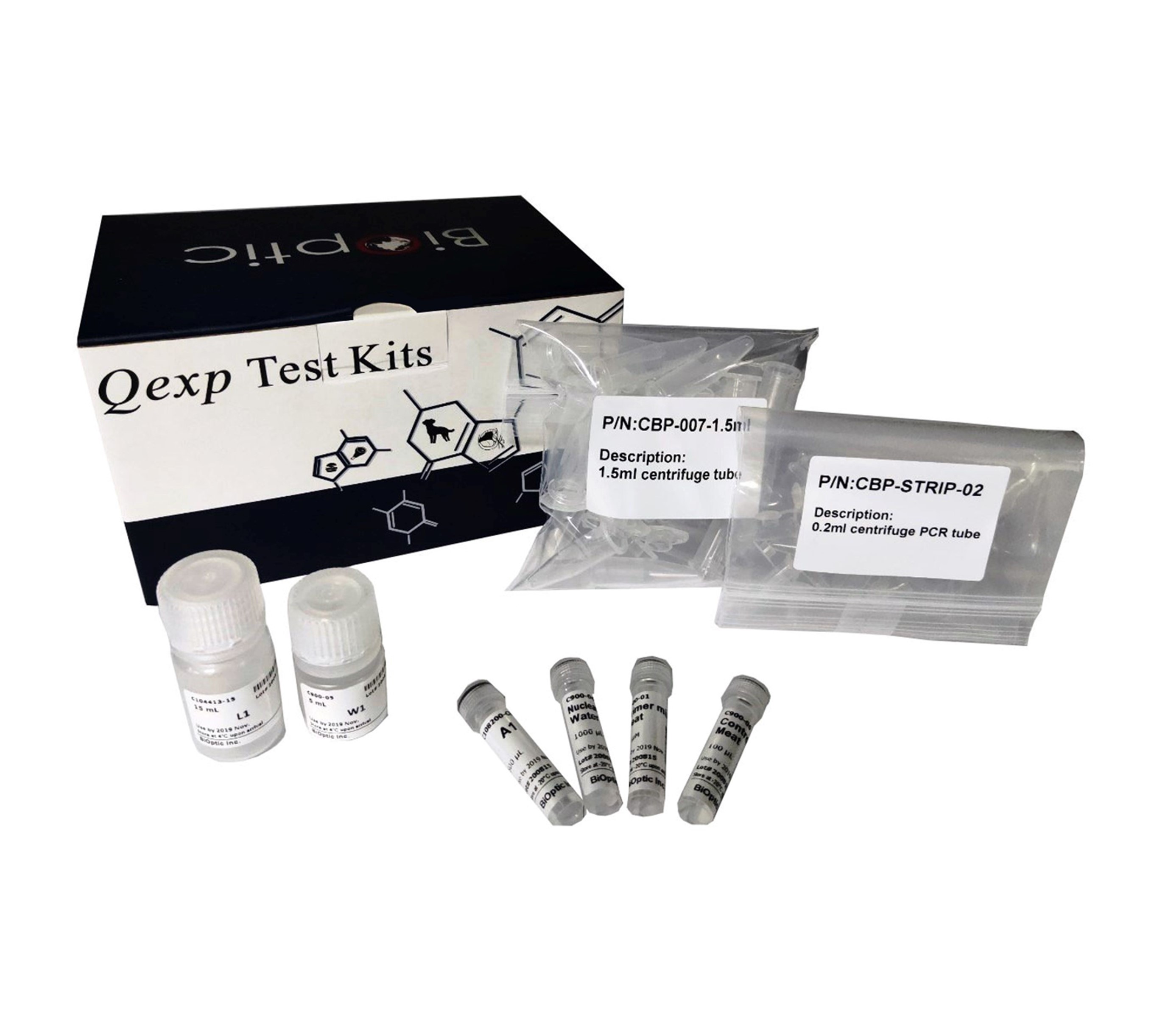
BiOptic Inc.
Leader in Innovative Capillary Electrophoresis Products.
BiOptic Inc. is a biotechnology instrumentation company that develops innovative scientific products for research and clinical applications.

High Resolution
DNA sample with 1 bp different can be resolved on Qsep Series platforms
LEARN MORE ›
CRISPR/Cas
CRISPR is a Genome Engineering technique that uses protein to interact with DNA. Genome editing begins with dsDNA cleavage. Cas9 is a dual-RNA-guided dsDNA endonuclease that is programmed by a single chimeric RNA. The guide RNA positions the Cas9 enzyme to the correct position on the target DNA for cleavage.
LEARN MORE ›
High Molecular Weight DNA (HMW DNA)
With ongoing innovation and improvements in the biotechnology industry now scientists are able to isolate longer and longer lengths of DNA at the high molecular weight (HMW DNA) for genome research. For downstream applications such as third-generation sequencing, the HMW DNA quality becomes an important issue.
LEARN MORE ›Recent News
Mar 28, 2024
Dec 21, 2023
Dec 18, 2023
Dec 14, 2023
Dec 07, 2023